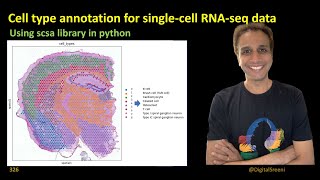
Single cell and spatial omics: A short introduction to the core concepts of scRNA-seq and more
ฝัง
- เผยแพร่เมื่อ 19 พ.ค. 2024
- A short introduction to the core concepts on single-cell omics data and spatial omics data. I will start by introducing how these types of data relate to and differ from normal omics data and concisely explain the typical experimental workflows used to produce such data. I will then talk about some of the computational aspects of analyzing single-cell data, including pooling of cells, clustering to identify cell types, and the concept of pseudo-time. Finally, I will briefly talk about how analysis of spatial omics data relate to analysis of images.
0:00 Introduction: reminder of omics is and introduction to single-cell and spatial data
0:42 Single-cell workflow: dissociated cell culture, isolation of single cells, omics on single cells, UMIs, and multiplexing
2:24 Spatial workflow: in situ capture, spatial indexing, spatial RNA-seq, single-cell resolution, and laser capture microdissection
3:48 Computational analysis: single cell vs. bulk data, cells vs. samples, pooling, cell-type clustering, pseudo-time reconstruction, and analysis of spatial patterns







I watched several videos trying to find an easy explanation about this. You made it so simple. Thank you. Best regards from México:)
Glad it was helpful!
Wow, such cutting-edge topics you are bringing to this channel. This is definitely one of my favorite areas of bioinformatics. There is so much more to say about scRNA-seq, but your video offers a nice and concise introduction. I very much agree that the sparsity of scRNA-seq data can be overcome by generating pseudo-bulks. In my case, I do clustering, followed by cell assignment and random pseudo-bulking. In this way, I artificially enhance the depth of my measuring units, while I can keep the cell type-specific information :)
Thanks a lot! What we're doing a lot is not to assign cell types, but rather to use variational autoencoders to compress the cells into a lower-dimensional latent space. It accomplishes much the same, since compressing the data into a lower-dimensional space inherently means that you are in some way averaging/summing up cells.
@@larsjuhljensen That sounds really interesting. I've been thinking along these lines too but in a less sophisticated way (just using knn to gather cells for pseudo-pooling across a lower dimensional space).
I would need to learn about autoencoders to really understand your strategy, but I kind of get the point.
You can read about it on bioRxiv: doi.org/10.1101/2022.07.06.499022
Or you can watch Mikaela from my group explain it: th-cam.com/video/XyfsK4oujVc/w-d-xo.html
That was amazing sir. You teaching bioinformatics and sharing your knowledge of omics, is the best course that one can take.
Great introduction thank you.
You are welcome!
wow, thank you so much.
You're welcome, I hope this short overview was useful!
how am I supposed to understand without even one illustration picture
I would have loved to have good, simple graphical illustrations of the many different assays too, but the figures I could find were overly complicated and not suitable for a short overview presentation.